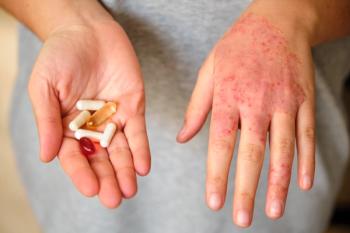
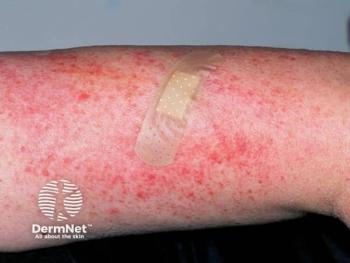
Atopic dermatitis associated with different bacteria depending on disease severity
NIH investigators find that Staphylococcus epidermidis is predominant in less severe cases, while Staphylococcus aureus is associated with patients who have more severe disease.
Using advanced, high-resolution sequencing techniques, investigators are gaining more detailed insights than ever before into the microbial communities associated with atopic dermatitis (AD), and how specific staphylococcal strains may contribute to the complexity of this condition.
Investigators at the National Institutes of Health (NIH) employed the approach, known as shotgun metagenomic sequence analysis, as part of an investigation of bacterial strain diversity in children with moderate to severe AD.
They found that AD flares are predominantly associated with Staphylococcus aureus in patients who have more severe disease, while Staphylococcus epidermidis is predominant in less severe cases. Sequencing analyses further demonstrated that clonal S. aureus strains were present in more severe patients, while heterogeneous S. epidermidis strains were present in all patients, regardless of disease severity. Results were
“Our findings are highlighting that there is much more heterogeneity in this disease, particularly with regard to the bacteria we are seeing on patients’ skin,” said Heidi H. Kong, M.D., M.H.Sc., Investigator, Dermatology Branch, National Institutes of Health, Bethesda, Md.
The relationship between inflammation in AD and skin bacteria - particularly S. aureus - is well known, and antimicrobial treatments do seem to at least reduce the abundance of these bacteria in the skin. Most studies to date have used traditional typing methods or sequencing of bacterial marker genes to study these bacteria in AD patients.
By contrast, shotgun metagenomic sequence analysis provides greater resolution that allows investigators to elucidate differences beyond species, all the way down to strain-level diversity, Dr. Kong said in an interview with Dermatology Times.
Shifts over time in bacterial community composition were noted in the NIH study, which included 11 children with moderate to severe AD along with 7 healthy children. By looking at samples at baseline (typical disease state), during flares, and after skin-directed treatment, they were able to show that worsening disease was inversely correlated with reduced skin bacterial diversity, particularly at the inner elbow and behind the knees, two predominant sites for AD (r = -0.58, P = 4.5 x 10-4).
While previous studies2 have shown a positive correlation between the course of AD and the presence of staphylococci, Dr. Kong and colleagues were able to look more closely at the relative abundance of specific species, including S. aureus, S. epidermidis, Staphylococcus hominis, and Staphylococcus capitis. Of those, only S. aureus had a significant change from flare to post-flare in mean relative abundance (from 28 +/- 8.8% to 2.3 +/- 0.8%, P < 0.014).
Moreover, the S. aureus strains demonstrated monoclonality in severe AD. Investigators used a previously validated strain-tracking approach for S. aureus and S. epiderimidis3,4 and for S. aureus, additionally mapped microbial reads against a database of 215 S. aureus genomes. They found the 5 patients with more severe disease were predominantly colonized by the bloom of a one particular strain. Notably, the strains of S. aureus these severe patients were colonized with were different, supporting previous investigations that suggest there is no single dominant S. aureus clone shared by all AD patients.
“The fact that the strain was different for each patient emphasizes the complexity of this disease. For each person, the development of AD is a multifactorial process,” Dr. Kong said. “Likely, genetics, the immune system, and skin barrier all play a role. So, why is there a specific strain in one person, and a different strain in another? Some of the future research may come down to a personalization of understanding what a specific S. aureus may be doing in each patient.”
After that, they topically colonized mice with staphylococcal strains they had isolated from these patients, as well as healthy controls. They noted some strain-specific differences in skin inflammation and immune response, and in particular, saw epidermal thickening and expansion of cutaneous T helper cells in mice colonized with S. aureus isolates from patients with more severe flares.
Clinical implications
The clinical implications of these findings are not clear, at least for now. While they provide greater insight into the diversity of staphylococcal strains that underlie AD, it is still not known, for example, whether or not S. aureus is triggering flares.
“If we can show that a particular strain is important in worsening AD for one particular person, then potentially, we could develop ways to target those particular strains. Unfortunately, if the strain is different for each person, it may be difficult to generalize these treatments, but we have a lot more that we need to explore.”
Next steps in this line of research, Dr. Kong said, will be to hone in on whether the particular strains identified in this study have a functional component contributing to AD.
“As we dig more into the research, we need to be very cognizant of what kinds of strains we are testing,” Dr. Kong said. “We should be cautious about taking an off-the-shelf strain to study. We may want to more carefully look at whether these are the strains that we can find on an AD patient.”
Summary
- Shotgun metagenomic sequence analysis was employed as part of an investigation of bacterial strain diversity in children with moderate to severe atopic dermatitis.
- Atopic dermatitis flares are predominantly associated with Staphylococcus aureus in patients who have more severe disease, while Staphylococcus epidermidis is predominant in less severe cases.
- Clonal S. aureus strains were present in more severe patients, while heterogeneous S. epidermidis strains were present in all patients.
- Findings highlight more heterogeneity in atopic dermatitis than previously known, though clinical implications are as yet unclear.
REFERENCES
1.) Byrd AL, Deming C, Cassidy SKB, Harrison OJ, Ng WI, Conlan S; NISC Comparative Sequencing Program, Belkaid Y, Segre JA, Kong HH. Staphylococcus aureus and Staphylococcus epidermidis strain diversity underlying pediatric atopic dermatitis. Sci Transl Med. 2017 Jul 5;9(397).
2.) Williams MR, Gallo RL. The role of the skin microbiome in atopic dermatitis. Curr Allergy Asthma Rep. 2015 Nov;15(11):65.
3.) Oh J, Byrd AL, Deming C, Conlan S; NISC Comparative Sequencing Program, Kong HH, Segre JA. Biogeography and individuality shape function in the human skin metagenome. Nature. 2014 Oct 2;514(7520):59-64.
4.) Oh J, Byrd AL, Park M; NISC Comparative Sequencing Program, Kong HH, Segre JA. Temporal Stability of the Human Skin Microbiome. Cell. 2016 May 5;165(4):854-66.