- Dermatology Times, August 2021 (Vol. 42, No. 8)
- Volume 42
- Issue 8
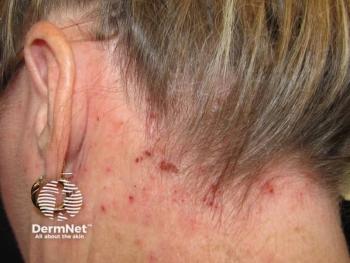
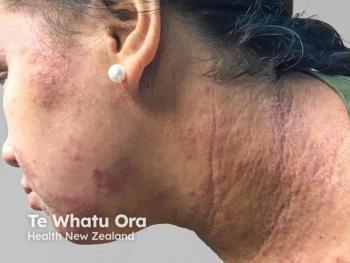
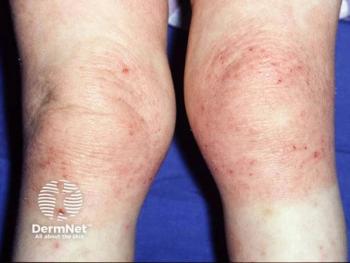
Analysis: Discrepancies Between ‘Atopic Dermatitis’ and ‘Eczema’
Investigators show that both terms tend to be used in different contexts within the biomedical literature.
Although the terms “eczema” and “atopic dermatitis” are typically used interchangeably, use of either term may have implications on meta-analyses, systematic reviews, and text-mining, according to a new study.
The study investigators found significant discrepancies between the terms in regard to the type of literature and information retrieved through database searches.
The team, led by Clément Frainay, PhD, School of Public Health, Imperial College London, searched the PubMed search engine for both terms, implementing machine learning to investigative the contexts in which either term has been used.
“We [chose] the corpus that pre-dates the recent 2017 recommendations on terminology, as their endorsement and impact cannot be properly assessed yet at time of writing.,” they noted.
Frainay and team then used a decision tree approach where they trained a model to predict whether an article would be indexed with eczema or atopic dermatitis tags. Text-mining tools were used to extract biological entities associated with either term.
Findings
Overall, the investigators found that an ‘AD’ query produced articles related to veterinary science, biochemistry, cellular, and molecular biology—while 'eczema' produced results linked to public health, infectious disease, and respiratory system.
Further, there were differences in ‘AD’ and ‘Eczema’-related terms from the Medical Subject Headings (MeSH) thesaurus, a resource that provides a controlled biomedical vocabulary. More specifically, there was a 52% agreement between the top 40 lists.
“The presence of terms related to cellular mechanisms, especially allergies and inflammation, characterized AD literature,” the investigators wrote. “The metabolites mentioned more frequently than expected in articles with AD tag differed from those indexed with eczema.”
They also found that eczema produced fewer enriched genes when compared with AD. Also, the genes retrieved from AD covered an average of 91.2% of the total number of genes retrieved from eczema or AD.
Gene Set to Diseases (GS2D) retrieved only 2 genes through an eczema search, which were also in the ‘AD’ search.
However, with Polysearch, 17 genes retrieved from an eczema search were not retrieved from an AD search.
Perspective
Frainay and team emphasized the differing contexts in which both terms had been used.
“Our results are an example of the implications of disease name ambiguity on text mining approaches, and emphasize the need to characterize, in terms of topics and content, the literature associated with each term and detect when two ‘synonymous’ disease names don’t carry the same information,” they wrote.
They suggested that a systematic approach should be employed using both terms jointly. Even more, they proposed decision tree learning as a tool to identify and characterize term ambiguity.
“Our results should raise awareness of the potential bias imputed to the term used when relying on text-mining approach and exemplify the importance of setting proper time frame and terms when querying publication database,” the investigators concluded.
The study, “
This article was originally published by our sister publication
Articles in this issue
over 4 years ago
Shrinking Staff Levels Can Lead to Unhappy Patientsover 4 years ago
Hair Hang-Upsover 4 years ago
Curcumin Shows Promise as New Sunscreen Filterover 4 years ago
Immunotherapy Helps Manage Skin Disease in Patients with Cancerover 4 years ago
Study: Curcumin Nanoparticles as a Photoprotective Adjuvantover 4 years ago
Novel Rosacea Treatment Demonstrates Efficacyover 4 years ago
Filler Hypersensitivity Post COVID-19 Illnessover 4 years ago
Climate Change Takes Toll on Skin Healthover 4 years ago
Ritlecitinib Delivers Topline Results in Alopecia Areata StudyNewsletter
Like what you’re reading? Subscribe to Dermatology Times for weekly updates on therapies, innovations, and real-world practice tips.