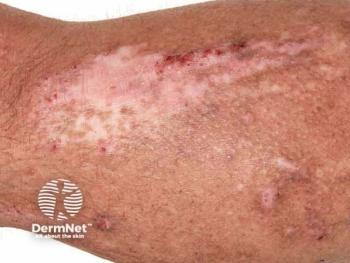
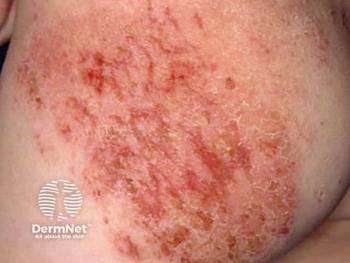
The factors that drive atopic dermatitis
Key Takeaways
- Genetic and epigenetic pathways are crucial in atopic dermatitis, with environmental factors increasing its prevalence.
- Filaggrin gene mutations and phthalate exposure are significant risk factors for atopic dermatitis.
There are two pathways that lead to atopic dermatitis. Their origins differ, but they both impair the immune system and skin barrier in AD.
A literature review by researchers at National Jewish Health in Denver shows that two major biologic pathways are responsible for driving the development of atopic dermatitis. One is genetic, the other epigenetic, but both can lead to changes in gene expressions that impairs the immune system and skin barrier.
Although it is, in part, a heritable disease, the prevalence of atopic dermatitis has risen in recent years due to environmental factors. The surge in new cases has led to more interest in epigenetics research which explains how lifestyle and air pollution can alter gene expressions leading to atopic dermatitis.
The last comprehensive review - published in 2009 - analyzed five linkage studies and 111 targeted gene association studies. In June 2009, the availability and appreciation of epigenetic data had just started to grow. Over the next several years, an explosion in technologies - including next-generation methods such as exome analysis and high-throughput genetic expression profiling - spurred the publication of more than 30 new studies between January 2015 and June 2016 alone.3
Earlier studies that focused on individual candidate genes helped establish a focus on mutations in the filaggrin gene (FLG) as the pre-eminent atopic dermatitis risk factor, write Lianghua Bin, M.D., Ph.D., and Donald Y. M. Leung, M.D., Ph.D., of the National Jewish Health in Denver.
The present review, published in a 2016 issue of the journal
A PubMed search yielded 19 candidate gene association studies (65 genes; with more than half having at least one positive association) and 13 genome-wide association studies (GWAS) in atopic dermatitis. Along with identifying additional associations and risk loci, Bin and Leung highlight how advances in genetic research have begun to illuminate the roles these defects play in the development of atopic dermatitis.
Barrier genes
Among skin barrier genes, FLG is the most significant risk factor for atopic dermatitis pathogenesis, and two of its known mutations - R501X and 2282del4 - have shown the strongest association for atopic dermatitis in Caucasians (18% and 48% of moderate to severe AD, respectively).4
Among environmental factors, recent research shows that phthalate metabolite levels are significantly associated with increased atopic dermatitis risk, and that children with FLG P478S mutations experience increased skin absorption of phthalate. Maternal FMG mutations, independent of mutation inheritance, also can act as strong environmental risk factors, the authors say.
Additional research implicates genes that impair formation of tight junctions within the epidermal granular layer. For example, owing to Th2 cytokines and genetic variants, expression of claudin-1/CLDN1 is reduced in AD-affected skin. Several studies also highlight the role of defects in the serine protease inhibitor of Kazal type 5 (SPINK5) gene, a crucial protease inhibitor for epidermal homeostasis.
Immunity genes
Regarding adaptive and innate immune response genes, the allergen sensitization, elevated serum IgE and increased expression of Type 2 cytokines associated with atopic dermatitis have long drawn attention to the Th2 axis. In this pathway, genetic variants in IL-4, the IL-4 receptor and IL-13 have shown positive associations with atopic dermatitis. Polymorphisms in IL-31, which is increased in atopic dermatitis lesions and serum and is involved in atopic dermatitis inflammatory responses, skin barrier dysregulation and severe itching, also have been reported.
Because thymic stromal lymphopoietin (TSLP)'s key function is to promote Th2 immune response, experts believe that TSLP plays an important role in atopic dermatitis pathogenesis. Both GWAS and targeted gene association studies have linked genetic variants in TSLP with AD risk, while expression of TSLP – which can be induced in epidermal epithelial cells by stimuli such as scratching and infections – is significantly increased in atopic dermatitis lesions. IL-33 and IL-25, which are also epithelium-derived promoters of the Th2 immune axis, appear to play key roles in atopic dermatitis pathophysiology, although not through genetic polymorphisms.
The increased prevalence of skin infections in AD suggests impairment in the innate immune system in AD-afflicted skin. Indeed, several pattern recognition receptor (PRR) signaling pathway genes including TLR2 and TLR9 have been associated with AD. Along with the direct effect of such genetic modifications, review authors write, "the attenuation of the normal antimicrobial response caused by overexpression of Th2 cytokines in skin is especially relevant in AD." Studies have shown, for instance, that Th2 cytokines can reduce the expression of human beta defensin 3 and LL-37 in epidermal keratinocytes.
Altogether, Bin and colleagues conclude, genetic variations and acquired impairment of innate immune responses may contribute to Th2 polarization in AD. However, they add, due to small sample sizes in candidate gene assays, the nascent nature of epigenetic research and the need to more thoroughly map genetic loci highlighted by GWAS, additional research is needed.
In summary
- There are 34 risk loci for atopic dermatitis suggesting a role for genes in immune responses and epidermal skin barrier in atopic dermatitis.
- Atopic dermatitis is associated with decreased gene expression of epidermal differentiation complex genes and elevated Th2 and Th17 genes.
- Hypomethylation of TSLP and FCER1G in atopic dermatitis were reported.
- MiR-155, which targets the immune suppressor CTLA-4, was found to be significantly over-expressed in infiltrating T cells in atopic dermatitis skin lesions.
DISCLOSURES
Dr. Bin is an advisory board member for Astellas Pharma Inc.
REFERENCES
1.) Barnes KC. An update on the genetics of atopic dermatitis: scratching the surface in 2009. J Allergy Clin Immunol. 2010;125(1):16-31.
2.) Esparza-Gordillo J, Weidinger S, Fölster-Holst R, et al. A common variant on chromosome 11q13 is associated with atopic dermatitis. Nat Genet. 2009;41(5):596-601.
3.) Bin L, Leung DY. Genetic and epigenetic studies of atopic dermatitis. Allergy Asthma Clin Immunol. 2016;12:52. eCollection 2016.
4.) O'Regan GM, Sandlands A, McLean WH, Irvine AD. Filaggrin in atopic dermatitis. J Allergy Clin Immunol. 2008;122(4):689-93.