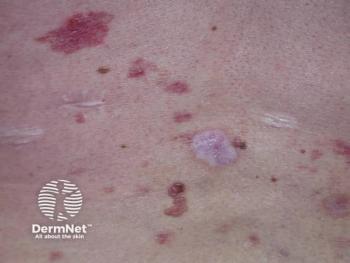
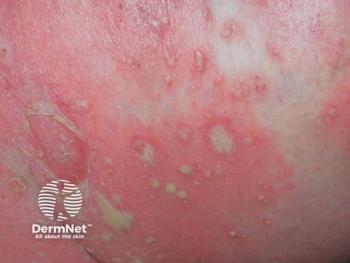
New psoriasis patient profile
Key Takeaways
- Metabolomics reveals a unique serum profile in psoriasis, with significant differences in amino acids, urea, and other metabolites compared to controls.
- Machine learning models, using metabolomic data, show promise in diagnosing and monitoring psoriasis, with 77% sensitivity and 74% specificity.
Metabolomics could provide new statistical methods to help monitor psoriasis disease and progression.
Advances in metabolomics are providing insights into psoriasis beyond what can be gleaned from genetic or immunologic studies.
In one recent development in this emerging field, investigators defined a unique serum profile of psoriasis based on specific metabolites. Building on those results, they developed some advanced statistical models that suggest metabolomics might one day be promising for the diagnosis, monitoring, and treatment of patients with this disease.
Results of the study, which were published recently in the Archives of Dermatological Research,1 show that individuals with psoriasis exhibit significantly different serum concentrations of specific amino acids, urea, acylcarnitines, phosphatidylcholines, phytol, and other metabolites compared with control subjects.
“While dermatologists are already doing an excellent job, understanding the metabolomics and biochemical background of psoriasis will definitely improve the treatment of their patients,” says investigator Aigar Ottas, M.Sc., chair of medical biochemistry with the Institute of Biomedicine and Translational Medicine, University of Tartu, Tartu, Estonia.
To date, most scientific investigations of psoriasis have been focused on the genetic background or immunologic aspects of the disease. What Mr. Ottas and other researchers like him hope to do is foster a broader understanding of the disease through metabolomics, an emerging field that focuses on the identification and measurement of metabolites such as amino acids, carbohydrates and carbohydrate derivatives, and lipids.
“Although the understanding of the genetic background of a disease is very important, it does not allow to monitor the current state of a disease nor its progression,” Mr. Ottas tells Dermatology Times. “This is something that metabolomics could really contribute in.”
Metabolites make up unique psoriasis profile
To help define a metabolomic profile for psoriasis, Mr. Ottas collected fasting blood samples from a total of 55 psoriasis patients and 51 age- and sex-matched controls.
In one portion of the study, they used a targeted approach to analyze concentrations of known metabolites. A total of 19 metabolites were identified that differed significantly between psoriasis patients and controls. For example, serum from psoriasis patients had lower concentrations of acylcarnitines, such as nonaylcarnitine (0.4 +/- 0.01 µM vs 0.5 +/- 0.01 µM; P = 0.002), and had higher concentrations of amino acids such as glutamate (92.85 +/- 66.43 µM vs 49.06 +/- 22.76 µM; P = 0.002), phenylalanine (82.91 +/- 18.96 µM vs 72.46 +/- 13.51 µM; P = 0.026), and ornithine (99.79 +/- 29.44 µM vs 82.28 +/- 20.85 µM; P = 0.011).
In a second portion of the study, investigators used an untargeted approach to discover other metabolites that might be implicated in psoriasis. They found a total of 22 metabolites with concentrations that varied significantly between serum of psoriasis patients and controls; of these, 12 metabolites could be identified; these included urea, taurine, and phytol, among others.
Many of the metabolites identified in the study are either part of the urea cycle or very closely related. Other metabolites had an exogenous (ie, food) origin.
“Since the metabolism of psoriasis patients seems to be altered it might be worth investigating if and how much diet affects the onset and progression of the disease,” Mr. Ottas says.
Next:
Machine learning
Researchers then used machine learning techniques to build models, based on metabolomic data, that would accurately discriminate between psoriasis patients and controls. The specific technique they used is known as the random forest algorithm, a method that is popular in bioinformatics. One of the models they described in their study report, based on 15 metabolites, had a 77% sensitivity and 74% specificity averaged across five repetitions.
Statistical models such as these could potentially be used to help diagnose the disease and monitor the treatment’s effectiveness.
“We are living in very exciting times where a lot of resources are directed into machine learning and artificial intelligence,” Mr. Ottas says. “Our hope is that one day, these types of statistical models help the doctors deal with the disease better, thus freeing their time for other important medical procedures.”
The hope is that these statistical models will be useful independent from visible symptoms, which would help to diagnose the disease in its earliest stages. “This would also enable non-dermatologists to diagnose the disease better, and perhaps even enable the patients to screen their progression themselves, like we have with diabetes currently,” Mr. Ottas adds.
Emerging field
Results from this study are the latest to provide metabolomic insights into psoriasis, though literature in this field is very limited.
In one of the key metabolomic studies of psoriasis to date, Kamleh and colleagues2 found changes in specific free-circulating amino acids, such as arginine, alanine, proline, and glutamate. They found that treatment with etanercept, the TNF-alpha receptor blocker, led to a normalization of the amino acid levels.
Another study of psoriasis metabolomics, published by Armstrong and colleagues,3 showed that patients with the disease had higher concentrations of glucuronic acid and alpha ketoglutaric acid, along with lower levels of glutamine and asparagine. They also found that, in comparison to patients with psoriasis alone, patients with both psoriasis and psoriatic arthritis had lower levels of alpha ketoglutaric acid and higher levels of lignoceric acid.
Now, these latest findings from Mr. Ottas and colleagues provide a snapshot of the metabolomic serum profile of psoriasis, along with some new statistical tools that could have utility in disease monitoring.
“The future is man and computers working together even more closely,” Mr. Ottas says. “Embracing this now will help to accelerate the onset of a medical age where doctors will have computer-based tools at their disposal which help the patients, reduce costs, and save time.”
Disclosures: Mr. Ottas reports no relevant conflicts of interest.
References
1. Ottas A, Fishman D, Okas TL, Kingo K, Soomets U. The metabolic analysis of psoriasis identifies the associated metabolites while providing computational models for the monitoring of the disease. 1. Arch Dermatol Res. 2017 Jul 10. doi: 10.1007/s00403-017-1760-1. [Epub ahead of print]
2. Armstrong AW, Wu J, Johnson MA, Grapov D., Azizi B, Dhillon J, Fiehn O. Metabolomics in psoriatic disease: pilot study reveals metabolite differences in psoriasis and psoriatic arthritis. F1000Res. 2014 Oct 21;3:248. doi: 10.12688/f1000research.4709.1. eCollection 2014.
3. Kamleh MA, Snowden SG, Grapov D, Blackburn GJ, Watson DG, Xu N, Ståhle M, Wheelock CE. LC-MS metabolomics of psoriasis patients reveals disease severity-dependent increases in circulating amino acids that are ameliorated by anti-TNFα treatment. J Proteome Res. 2015 Jan 2;14(1):557-66. doi: 10.1021/pr500782g. Epub 2014 Nov 12.