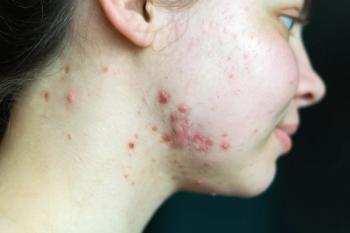
Knowledge of skin microbiome provides insights into dermatological conditions
Direct sequencing data, together with culture data, supplies researchers with information about communities of bacteria and fungi on the skin.
Research into the skin microbiome has revealed much diversity in bacteria and fungi that reside on the skin and may produce a greater understanding of skin conditions like atopic dermatitis (AD), according to an Investigator in the Dermatology Branch of the Center for Cancer Research, National Cancer Institute, National Institutes of Health in Bethesda, MD.
Speaking about the skin microbiome in health and disease at the annual meeting of the Canadian Dermatology Association, Heidi H. Kong, MD, MHSc, described how in healthy individuals, investigators aim to understand the composition of the skin microbiome and observe the stability and function of the skin microbes. On the other hand, in individuals with skin diseases, investigators try to grasp how the microbes promote the development of a condition like atopic dermatitis and how defects in skin immunity affect microbes.
"Humans are host to many microbes such as bacteria, fungi, and viruses," said Dr. Kong. "As clinicians, when we think of microbes, we often think of them as pathogenic, but microbes perform beneficial functions that are important for health, such as aiding in digestion."
Dr. Kong and co-investigators examined the skin microbiome in 12 children with moderate-to-severe eczema as well as 11 children who served as controls. She noted that Staphylococcus aureus colonization of the skin has been associated with the development of AD. Researchers performed 16S ribosomal RNA bacterial gene sequencing to better appreciate microbial communities and their potential role in the etiology of AD.1
Investigators found not only did S. aureus rise significantly during AD flares, but that, surprisingly, Staphylococcus epidermidis rose significantly during diseases flares, and that post therapy, there were rises in Streptococcus, Propionibacterium, and Corynebacterium.
While culture-based methods have shed light on human fungal skin diseases, direct sequencing data, in conjunction with culture data, provides more robust information about fungal and bacterial communities on the skin, noted Dr. Kong. Culture-based methods are less expensive than genetic sequencing, but genomic methods are dropping in cost, and they help detect microbes that cannot be easily grown, added Dr. Kong.
In one investigation, Dr. Kong and collaborators sequenced and analyzed fungal communities of 14 skin sites in 10 healthy adults. They determined that 11 core-body and arm sites were populated mainly by the fungi of the genus Malassezia. By contrast, they found that three foot sites consisting of plantar heel, toenail, and toe web demonstrated abundant diversity of fungi.
Of note in that investigation, one healthy volunteer had a more significantly diverse fungal profile than other subjects, and investigators discovered that subject completed a two-month course of oral antifungals seven months before study entry.1
"We observed that Malassezia predominate," said Dr. Kong. "We also determined that the foot site demonstrates greater fungal diversity. We also looked at the differences between bacteria and fungi."
Meta-genomic shotgun sequencing can help investigators to not just categorize the microbes that reside on the skin but also begin to understand the role of the microbes as they relate to the host, said Dr. Kong.
Various theories have been put forth for why individuals with AD experience the "atopic march" and develop other conditions like asthma and hay fever, and Dr. Kong noted that many are pursuing research avenues to understand the triggers of disease progression.
Another research effort involves comparing the skin microbiome in patients who have a dermatologic condition like AD to patients with primary immunodeficiencies who have AD-like skin disease, according to Dr. Kong, noting there is a need for host-skin response studies.
"We wanted to see if these diseases share the same microbiome features as atopic dermatitis," said Dr. Kong.
Investigators studied these patients who can suffer candidal infections and secondary aspergillus lung infections, said Dr. Kong.2
"We found that these patients harbor unique bacteria," said Dr. Kong. "We saw a greater prevalence of fungi, like aspergillus, in the nose and on the arms of these patients. Something in their immune system may not be able to rid their skin of these fungi which can be from the environment. These fungi are potentially putting them at risk for secondary aspergillus lung infections."
Research is focused on understanding microbes on the skin and how they can be influenced in skin diseases to bring skin back to a healthy state, said Dr. Kong.
Dr. Kong had no relevant disclosures.
Genomic Research. 2012 May;22(5):850-9.
Nature. 2013 Jun 20;498(7454):367-70.
Genomic Research. 2013 Dec;23(12):2103-14.