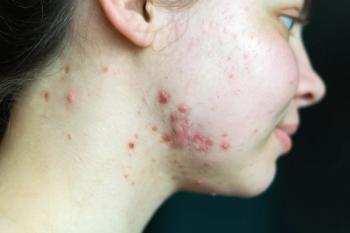
Link Between Gene Variants and Hidradenitis Suppurativa Identified
Researchers have isolated 2 genes, SOX9 and KLF5, that play a role in the development of hidradenitis suppurativa.
Researchers conducted a genome-wide association study (GWAS) to determine if genetic variants contribute to risk of developing hidradenitis suppurativa (HS) and found a correlation with 2 genes, SOX9 and KLF5. “The location of HS-associated variants in skin regulatory elements and the functions of nearby genes in HS disease pathways suggest that the variants alter expression of nearby genes to affect disease risk.”1
Patients enrolled in the HS Program for Research and Care Excellence (HS ProCARE) at the University of North Carolina (UNC) Department of Dermatology from August 2018 to July 2021 were recruited. After quality control, 720 patients were identified, with a mean age of 20.3 (10.57) for the onset of symptoms, 50% were Black, and 79.7% were female.
Samples from participants were obtained using Oragene_Dx saliva collection kits (DNA Genotek). Samples were analyzed for quality, quantity, and were genotyped at UNC facilities.
Researchers used information from the National Longitudinal Study of Adolescent to Adult Health (Add Health) for controls and performed liftover from hg19 to hg38. They used variants common to HS ProCARE and Add Health to perform genotype imputation and separated participants into 4 groups: European-ancestry HS cases, European-ancestry controls, African-ancestry HS cases, and African-ancestry controls and only kept variants with R2 of .08 or higher.
Study authors combined their results with samples from the UK Biobank (UKB), FinnGen (Finland), and BioVU (Vanderbilt University Medical Center).
Results found 2 genes, SOX9 and KLF5, to have variants. SOX9 maintains the structure of the hair follicle and assigns “stem cells to become, or differentiate into, the cells that line the hair follicle.2” It also activates 3 other genes, MMP1, MMP2, and IL-8, which are linked to nonmelanoma skin cancer development and inflammation.
In mouse models, overactivation of the KLF5 gene led to hyperkeratosis, blocked follicles, and skin erosions. An absence of KLF5 interferes with barrier function and has been associated with atopic dermatitis, non-healing wounds, and IL-17-driven colitis in mice.
The roles of SOX9 and KLF5 appear to be either in direct opposition or complementary to roles in epithelial differentiation.“ KLF5 and SOX9 expression demarcate epidermal and hair follicle differentiation, respectively.1”
Researchers also looked at comorbidities of HS and found a slightly significant link (P < .05) with inflammatory bowel disease, psoriasis, and type 2 diabetes.
Authors conclude that variants in the genes SOX9 and KLF5 could affect HS development because of their roles in “hair follicle and epidermal differentiation and inflammation.1”
References
- Sun Q, Broadaway KA, Edmiston SN, et al. Genetic variants associated with hidradenitis suppurativa. JAMA Dermatol. Published online July 26, 2023. Accessed July 27, 2023. doi:10.1001/jamadermatol.2023.2217
https://jamanetwork.com/journals/jamadermatology/fullarticle/2807701?guestAccessKey=eeef4297-5c4b-420b-9cc1-a74203bbe0e4&utm_source=silverchair&utm_medium=email&utm_campaign=article_alert-jamadermatology&utm_content=olf&utm_term=072623 - Researchers discover genetic locations for increased risk of hidradenitis suppurativa. News release. UNC Health. July 26, 2023. Accessed July 27, 2023.
https://news.unchealthcare.org/2023/07/researchers-discover-genetic-locations-for-increased-risk-of-hidradenitis-suppurativa/