Genetic Variants Associated With HS Identified
Genetic variants associated with hidradenitis suppurativa (HS) were identified, although a causal effect is not proven.
Common variants associated with hidradenitis suppurativa (HS) located near the SOX9 and KLF5 genes were linked with HS risk, and these or other neighboring genes may be associated with genetic disease risk and the development of clinical features like cysts, comedones, and inflammatory tunnels that are unique to HS, according to
The genetic understanding of HS is limited, and few genome-wide association studies (GWASs) have been conducted for HS; existing studies have not recognized significant risk loci.
“In this genetic association study, as with all GWASs, identifying variant associations near candidate genes with plausible roles in disease biology does not prove a causal effect of these variants or genes on disease risk,” said the researchers.
Researchers aimed to identify genetic variants associated with HS and to illuminate the underlying genes and genetic mechanisms.
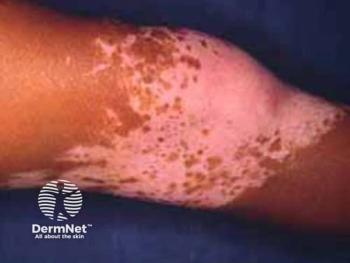
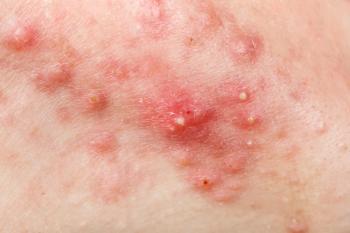
HS is a common and seriously morbid chronic inflammatory skin disease that is described as highly heritable. It’s characterized by painful nodules, abscesses, and tunnels with predilection for intertriginous sites. Siblings of affected patients have an almost 20-fold risk of developing HS.
First, a total of 753 patients with HS in the HS Program for Research and Care Excellence (HS ProCARE) at the University of North Carolina Department of Dermatology were recruited for this genetic association study from August 2018 to July 2021.
A GWAS was started for 720 patients (after quality control) with controls from the Add Health study and then meta-analyzed with 2 large biobanks, UK Biobank (247 cases) and FinnGen (673 cases). Variants at 3 loci were tested for replication in the BioVU biobank (290 cases), and data analysis was executed from September 2021 to December 2022.
The main outcome measures were loci identified, with association of P < 1×10–8 deemed significant.
A total of 720 patients were part of the analysis. Mean (SD) age at symptom onset was 20.3 (10.57) years and at enrollment was 35.3 (13.52) years; 360 patients were Black, and 575 were female. In a meta-analysis of the 4 studies, 2 HS-associated loci were pinpointed and replicated, with lead variants rs10512572 (P = 2.3×10–11) and rs17090189 (P = 2.1×10–8) near the SOX9 and KLF5 genes, respectively. Variants at these loci are found in enhancer regulatory elements identified in skin tissue.
The HS candidate gene SOX9, which encodes SRY-box transcription factor 9, is greatly expressed in human epidermis and the outer root sheath of hair follicles and is vital for follicular differentiation and maintenance. The SOX9 gene upregulates MMP1, MMP2, and IL-8, which are associated with tumorigenesis and inflammation. In human colitis, decreased intestinal KLF5 expression is associated with increased disease severity.
New insights into disease pathogenesis related to these genes might help estimate disease progression and new future treatment approaches. Additionally, genetic correlations of HS with common comorbidities like psoriasis, inflammatory bowel disease, and type 2 diabetes imply that shared genetic mechanisms might predispose to risk. Researchers recommend that further investigations are needed with better-powered HS GWASs to review other comorbidities and to reveal the mechanisms of shared risk.
Researchers also mentioned that functional studies need to review whether the HS-associated variants and surrounding regulatory elements can change SOX9 and KLF5 function or expression and, subsequently, affect the pathology of HS. These genes are strong candidates because they affect hair follicle and epidermal differentiation and inflammation. Other neighboring genes like KLF12 could also play a role. Larger sample sizes and sequencing are needed to identify rare gene variants, like γ-secretase complex variants, that are implicated in small numbers of patients.
This study has some limitations. Some Add Health participants might have had HS, which would lead to misclassification bias toward the null. While HS ProCARE included patients with ancestry from Europe and Africa, small sample sizes limited the researchers’ ability to identify significant loci within ancestry groups or evaluate generalizability in other ancestries.
“Ultimately, understanding how genes lead to tunnel formation and an overactive inflammatory response may lead to therapeutic breakthroughs that target HS pathogenesis,” concluded the researchers.
Reference
Sun Q, Broadaway KA, Edmiston SN, et al. Genetic variants associated with hidradenitis suppurativa. JAMA Dermatol. Published online July 26, 2023. doi:10.1001/jamadermatol.2023.2217
[This article was originally published by our sister brand,
Newsletter
Like what you’re reading? Subscribe to Dermatology Times for weekly updates on therapies, innovations, and real-world practice tips.